08 Dictionaries
question Questionsobjectives Objectives
- How to associate two values with each other in one data structure?
- Understand the difference between dictionaries and other data structures.
- Use Dictionaries to combine sets of values and store different data structures within a dictionary.
- Give an example of how dictionaries are used in bioinformatics (BioPython).
time Time estimation: 20 minutes
8.1 Introduction
So far we’ve seen variables that store one value or a series of values (see section 5: lists, tuples and sets). There is another way of storing information where you associate one value with another value; in Python this is called a dictionary. Dictionaries provide a very useful way of quickly connecting different values to each other.
8.2 Dictionary creation & usage
It is best to think of a dictionary as a set of key:value pairs, with the requirement that the keys are unique (within one dictionary). Dictionaries are initiated by using curly brackets {}, and each pair of key:values is separated with a comma. This is how a dictionary would look like:
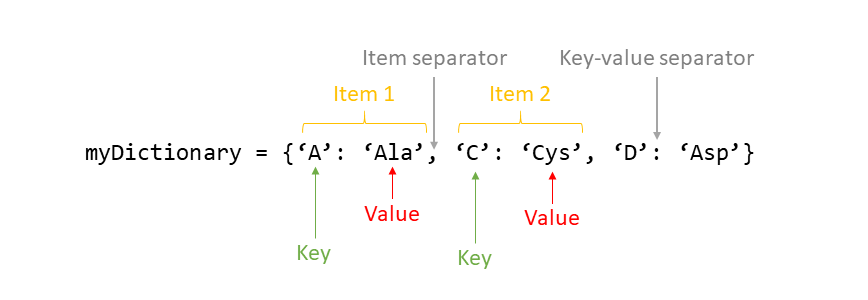
myDictionary = {'A': 'Ala', 'C': 'Cys', 'D': 'Asp'}
myDictionary
You can recall values by using square brackets [ ] with the name of the key, or use the get()-method.
myDictionary['A']
myDictionary.get('C')
If you would like to add a new pair of key-value:
myDictionary['E'] = 'Glu'
myDictionary
Note however that keys are unique and if you try to add a key:value-pair with a key that already exists in the dictionary and a different value, it will overwrite the value.
myDictionary['A'] = 'Glu'
myDictionary
So keys are unique, values are not!
Dictionaries, like lists, have several useful built-in methods. The most frequently used are listed here below:
keys()to list the dictionary’s keysvalues()to list the values in the dictionaryget()call the value of a specified keypop()to remove the specified key and its values
Listing the keys within a dictionary:
myDictionary = {'A': 'Ala', 'C': 'Cys', 'D': 'Asp', 'E': 'Glu'}
myDictionary.keys()
Python tells us that the list is still in a dictionary-keys data structure type. If you would like to extract the keys for further processing, it’s probably better to transform them into a list:
list(myDictionary.keys())
Similarly for the values of a dictionary:
list(myDictionary.values())
We’ve already exploited the get method, with pop we can remove a key-value pair:
myDictionary.pop('E')
myDictionary
If you try to access a key that doesn’t exist, Python will give an error:
myDictionary = {'A': 'Ala', 'C': 'Cys', 'D': 'Asp', 'E': 'Glu'}
myDictionary['B']
You should therefore always check whether a key exists:
# Newlines don't matter when initialising a dictionary...
myDictionary = {
'A': 'Ala',
'C': 'Cys',
'D': 'Asp',
'E': 'Glu',
'F': 'Phe',
'G': 'Gly',
'H': 'His',
'I': 'Ile',
'K': 'Lys',
'L': 'Leu',
'M': 'Met',
'N': 'Asn',
'P': 'Pro',
'Q': 'Gln',
'R': 'Arg',
'S': 'Ser',
'T': 'Thr',
'V': 'Val',
'W': 'Trp',
'Y': 'Tyr'}
if 'B' in myDictionary.keys():
print(myDictionary['B'])
else:
print("myDictionary doesn't have key 'B'!")
However, it’s much cleaner if you use the get() method as it doesn’t return an explicit error if a key doesn’t exist in your dictionary. Instead it will return a None-value.
type(myDictionary.get('B'))
hands_on Exercise 8.2.1
Use a dictionary to track how many times each amino acid code appears in the following sequence:
SFTMHGTPVVNQVKVLTESNRISHHKILAIVGTAESNSEHPLGTAITKYCKQELDTETLGTCIDFQVVPGCGISCKVTNIEGLLHKNNWNIED NNIKNASLVQIDASNEQSSTSSSMIIDAQISNALNAQQYKVLIGNREWMIRNGLVINNDVNDFMTEHERKGRTAVLVAVDDELCGLIAIADTTip: use the one-letter code as key in the dictionary, and the count as value.
solution Solution
# Use a dictionary to track how many times each amino acid code appears in the following sequence: # SFTMHGTPVVNQVKVLTESNRISHHKILAIVGTAESNSEHPLGTAITKYCKQELDTETLGTCIDFQVVPGCGISCKVTNIEGLLHKNNWNIEDNNIKNASLVQIDASNEQSSTSSSMIIDAQISNALNAQQYKVLIGNREWMIRNGLVINNDVNDFMTEHERKGRTAVLVAVDDELCGLIAIADT # Tip: use the one-letter code as key in the dictionary, and the count as value. mySequence = "SFTMHGTPVVNQVKVLTESNRISHHKILAIVGTAESNSEHPLGTAITKYCKQELDTETLGTCIDFQVVPGCGISCKVTNIEGLLHKNNWNIEDNNIKNASLVQIDASNEQSSTSSSMIIDAQISNALNAQQYKVLIGNREWMIRNGLVINNDVNDFMTEHERKGRTAVLVAVDDELCGLIAIADT" # First way to do this, using sets (condensed) aminoAcidCount = {} myUniqueAminoAcids = set(mySequence) for aaCode in myUniqueAminoAcids: print("Amino acid {} occurs {} times.".format(aaCode,mySequence.count(aaCode))) aminoAcidCount[aaCode] = mySequence.count(aaCode)solution Solution
# Another way to do this, a little bit more elaborate and using the myDictionary as a reference for iteration mySequence = "SFTMHGTPVVNQVKVLTESNRISHHKILAIVGTAESNSEHPLGTAITKYCKQELDTETLGTCIDFQVVPGCGISCKVTNIEGLLHKNNWNIEDNNIKNASLVQIDASNEQSSTSSSMIIDAQISNALNAQQYKVLIGNREWMIRNGLVINNDVNDFMTEHERKGRTAVLVAVDDELCGLIAIADT" myDictionary = { 'A': 'Ala', 'C': 'Cys', 'D': 'Asp', 'E': 'Glu', 'F': 'Phe', 'G': 'Gly', 'H': 'His', 'I': 'Ile', 'K': 'Lys', 'L': 'Leu', 'M': 'Met', 'N': 'Asn', 'P': 'Pro', 'Q': 'Gln', 'R': 'Arg', 'S': 'Ser', 'T': 'Thr', 'V': 'Val', 'W': 'Trp', 'Y': 'Tyr'} lengthDict = len(myDictionary.keys()) for aa in range(lengthDict): aaCode = list(myDictionary.keys())[aa] aaCount = mySequence.count(aaCode) print("Amino acid {} occurs {} times.".format(aaCode,aaCount))
8.3 A practical example of dictionaries
An practical example of dictionaries can be found in Biopython. Imagine that we want to extract some information from a GenBank file (NC_005816)
# Imports the SeqIO object from Biopython
from Bio import SeqIO
# Reads in (just one record of) the GenBank file
record = SeqIO.read("data/NC_005816.gb","genbank")
print(record)
The SeqRecord object (which we see here) has an id, name and description as well as a sequence. For other (miscellaneous) annotations, the SeqRecord object has a dictionary attribute annotations. Most of the annotations information gets recorded in the annotations dictionary.
print(record.id)
print(record.name)
print(record.description)
#print(record.seq)
record.annotations
record.annotations['organism']
record.annotations['source']
(In general, organism is used for the scientific name (in Latin, e.g. Arabidopsis thaliana), while source
will often be the common name (e.g. thale cress). In this example, as is often the case, the two fields are
identical.)
record.annotations['accessions'] # This could be a list of values, hence the list.
8.4 More with dictionaries
As mentioned here above, the value associated with a key can consist of a list with values (instead of one single value). In the example below we save the information of an experiment in a dictionary. The key that saves the date information contains a list of fictive dates (01-01-2020 and 02-01-2020):
TriplicateExp1 = {'name': 'experiment 1', 'pH': 5.6, 'temperature': 288.0, 'volume': 200, 'calibration':'cal1', 'date':['01-01-2020','02-01-2020']}
TriplicateExp1
For the keys, however, the data structures should be immutable (so tuples are OK, lists are not). Recall that keys have to be unique; if you add a key that already exists, the old entry will be overwritten:
dates = ('date1','date2') # tuple
TriplicateExp1[dates] = ['01-01-2020','02-01-2020']
TriplicateExp1
It is also possible to have a so-called nested dictionary, in which there is a dictionary within a dictionary. Here we make two more dictionaries with information about the triplicate experiment. The information of each experiment is thus assembled in a separate dictionary. Then, the three dictionaries are combined into one dictionary.
TriplicateExp2 = {'name': 'experiment 2', 'pH': 5.8, 'temperature': 286.0, 'volume': 200, 'calibration':'cal1', 'date':'03-01-2020'}
TriplicateExp3 = {'name': 'experiment 3', 'pH': 5.4, 'temperature': 287.0, 'volume': 200, 'calibration':'cal1', 'date':'04-01-2020'}
Triplicate = {
'exp1':TriplicateExp1,
'exp2':TriplicateExp2,
'exp3':TriplicateExp3
}
Triplicate
keypoints Key points
- We learned why it can be beneficial to store information in dictionaries
- Dictionaries can be nested within each other and can contain multiple different data structures.
Useful literature
Further information, including links to documentation and original publications, regarding the tools, analysis techniques and the interpretation of results described in this tutorial can be found here.
